Electrostatics Benchmarks#
This page presents benchmark results for electrostatic interaction methods including Ewald summation, Particle Mesh Ewald (PME), and Damped Shifted Force (DSF) across different GPU hardware. Results show the scaling behavior with increasing system size for both single-system and batched computations.
Warning
These results are intended to be indicative only: your actual performance may vary depending on the atomic system topology, software and hardware configuration and we encourage users to benchmark on their own systems of interest.
How to Read These Charts#
Time Scaling : Median execution time (ms) vs. system size. Lower is better. Timings include both real-space and reciprocal-space contributions when running “full” mode.
Throughput : Atoms processed per millisecond. Higher is better. This indicates the scaling point where the GPU saturates.
Memory : Peak GPU memory usage (MB) vs. system size. This is particularly useful for estimating/gauging memory requirements for your system.
Ewald Summation#
The Ewald summation method splits the Coulomb interaction into real-space and reciprocal-space components. This is the traditional \(O(N^{3/2})\) to \(O(N^2)\) method depending on parameter choices.
Simple comparison of single (non-batched) system computations between backends, where we scale up the size of the supercell.
Time Scaling
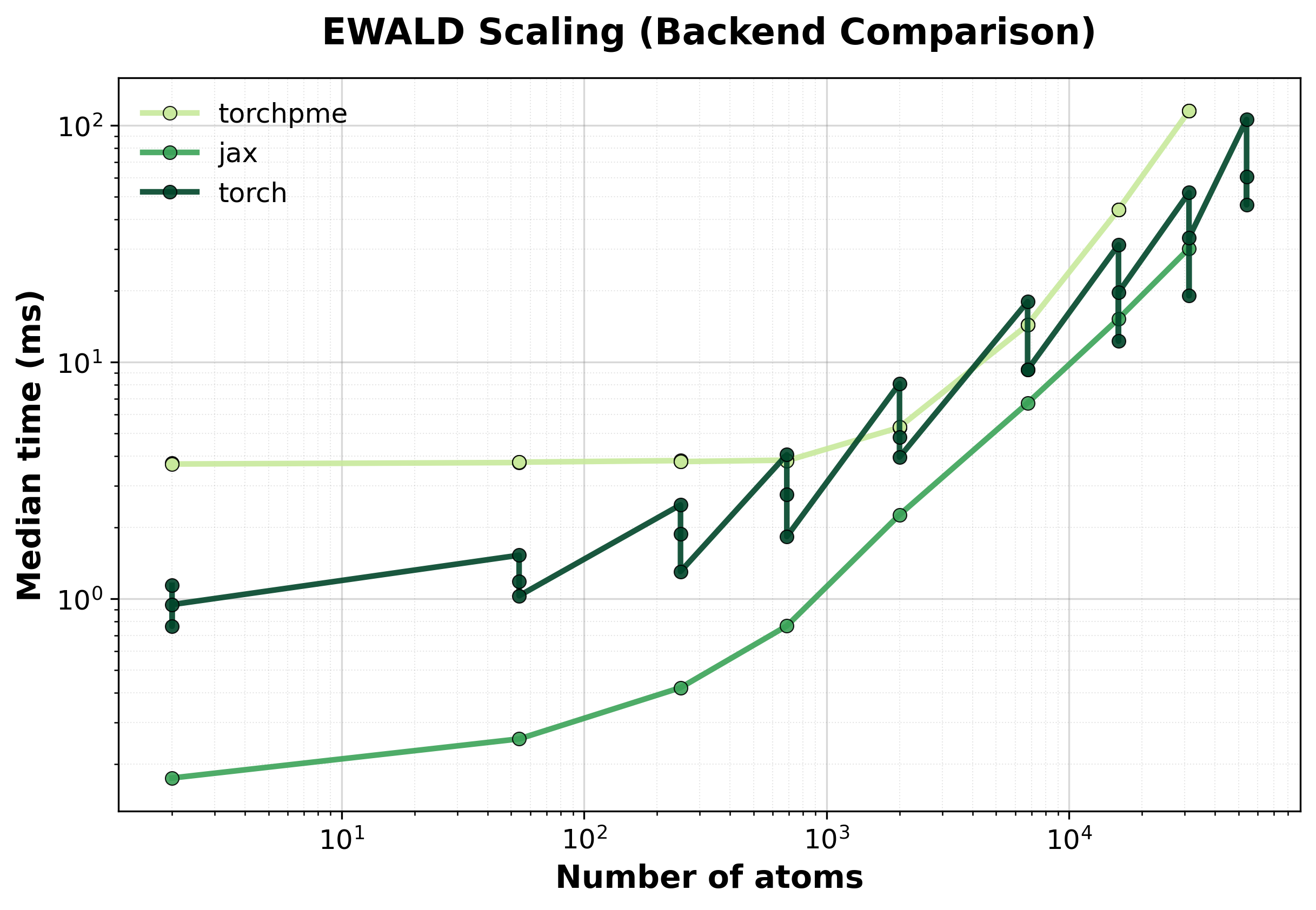
Median execution time comparison between backends for single systems.#
Throughput
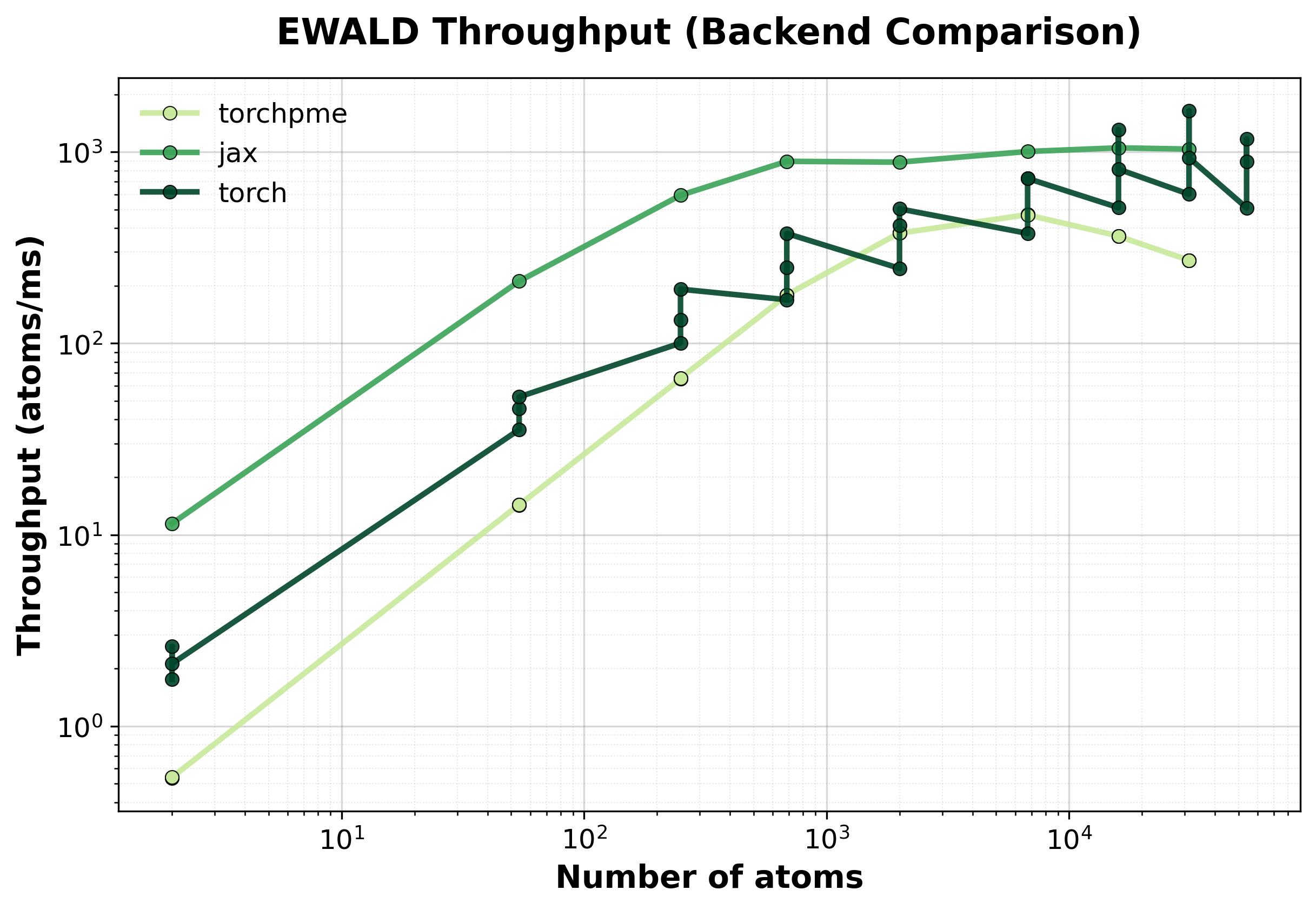
Throughput (atoms/ms) comparison between backends. Higher values indicate better performance.#
Memory Usage
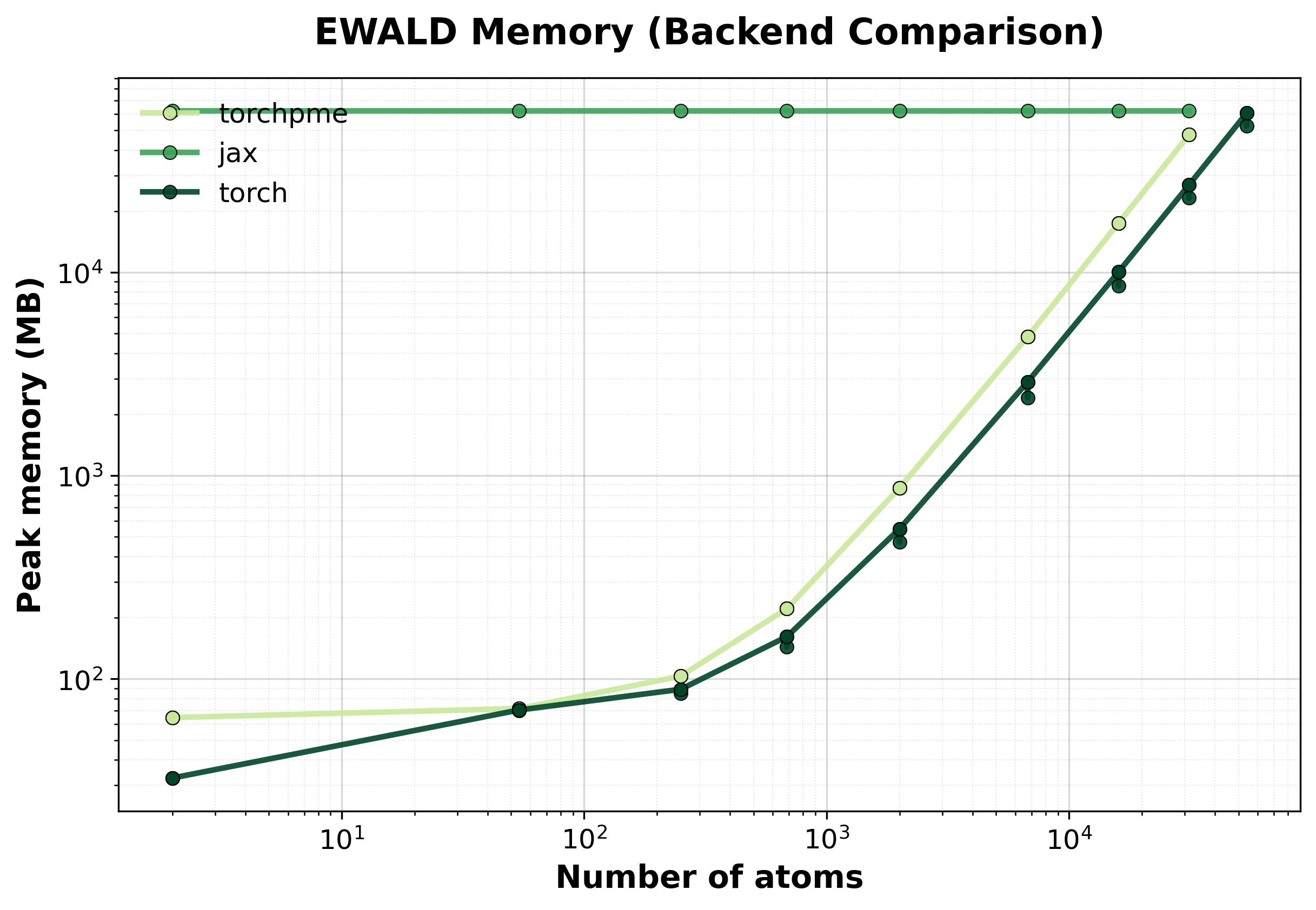
Peak GPU memory consumption comparison between backends. Lower is better, indicating that the backend has lower memory requirements.#
Scaling of single and batched Ewald computation with the nvalchemiops Torch backend.
Shows how performance scales with different batch sizes.
Time Scaling
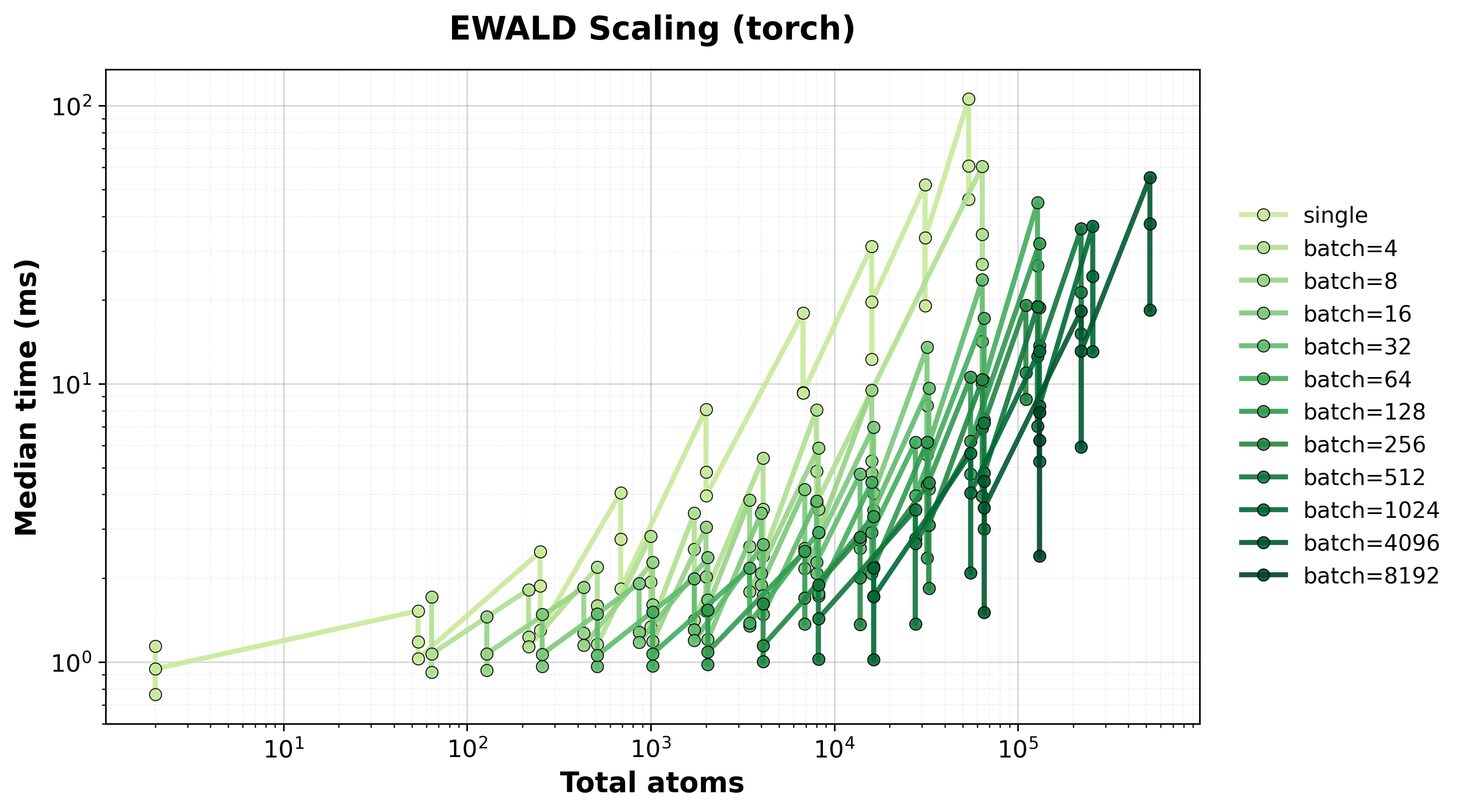
Execution time scaling for single and batched systems.#
Throughput
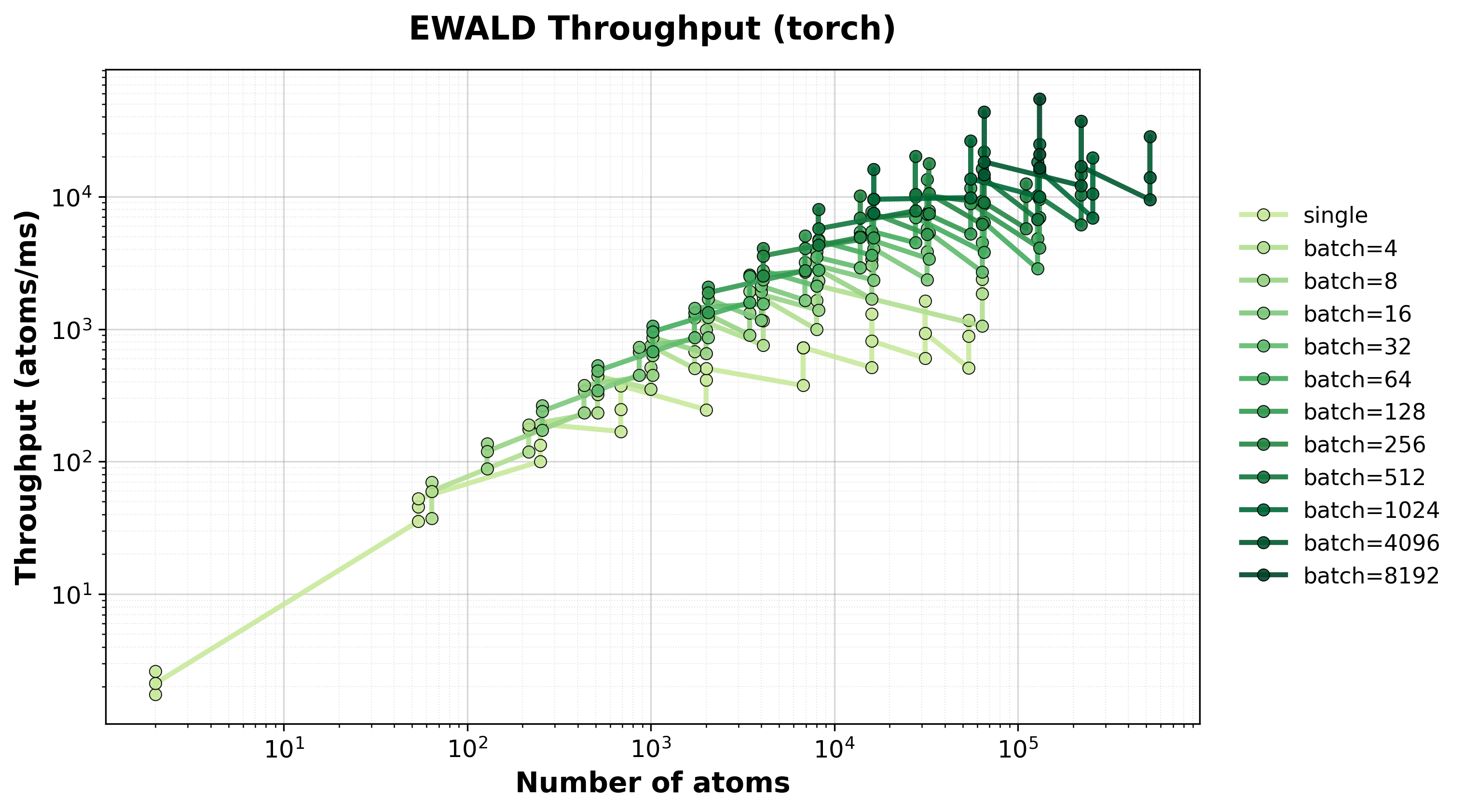
Throughput (atoms/ms) for single and batched systems.#
Memory Usage
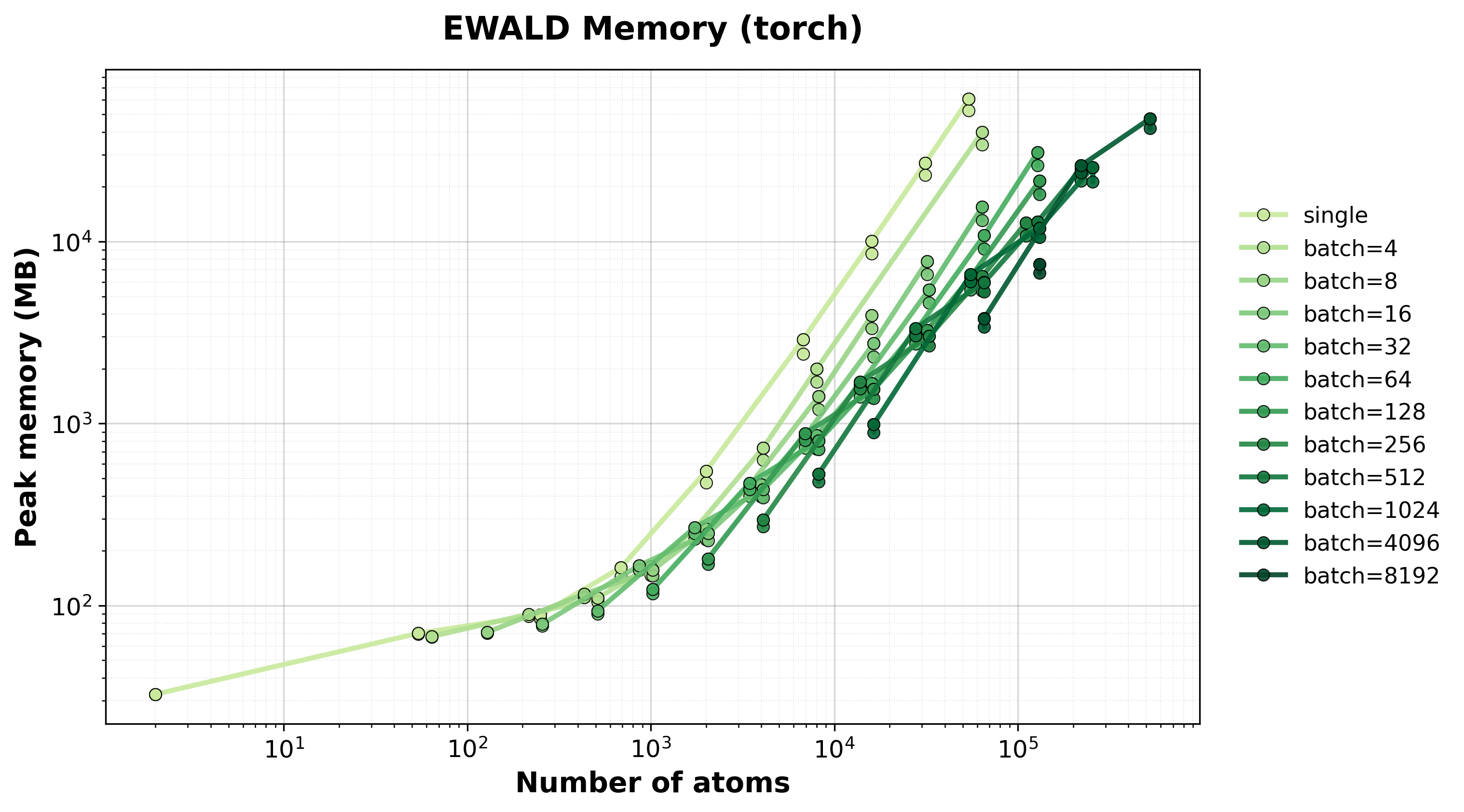
Peak GPU memory consumption for single and batched systems.#
Scaling of single and batched Ewald computation with the nvalchemiops JAX backend.
Shows how performance scales with different batch sizes.
Time Scaling
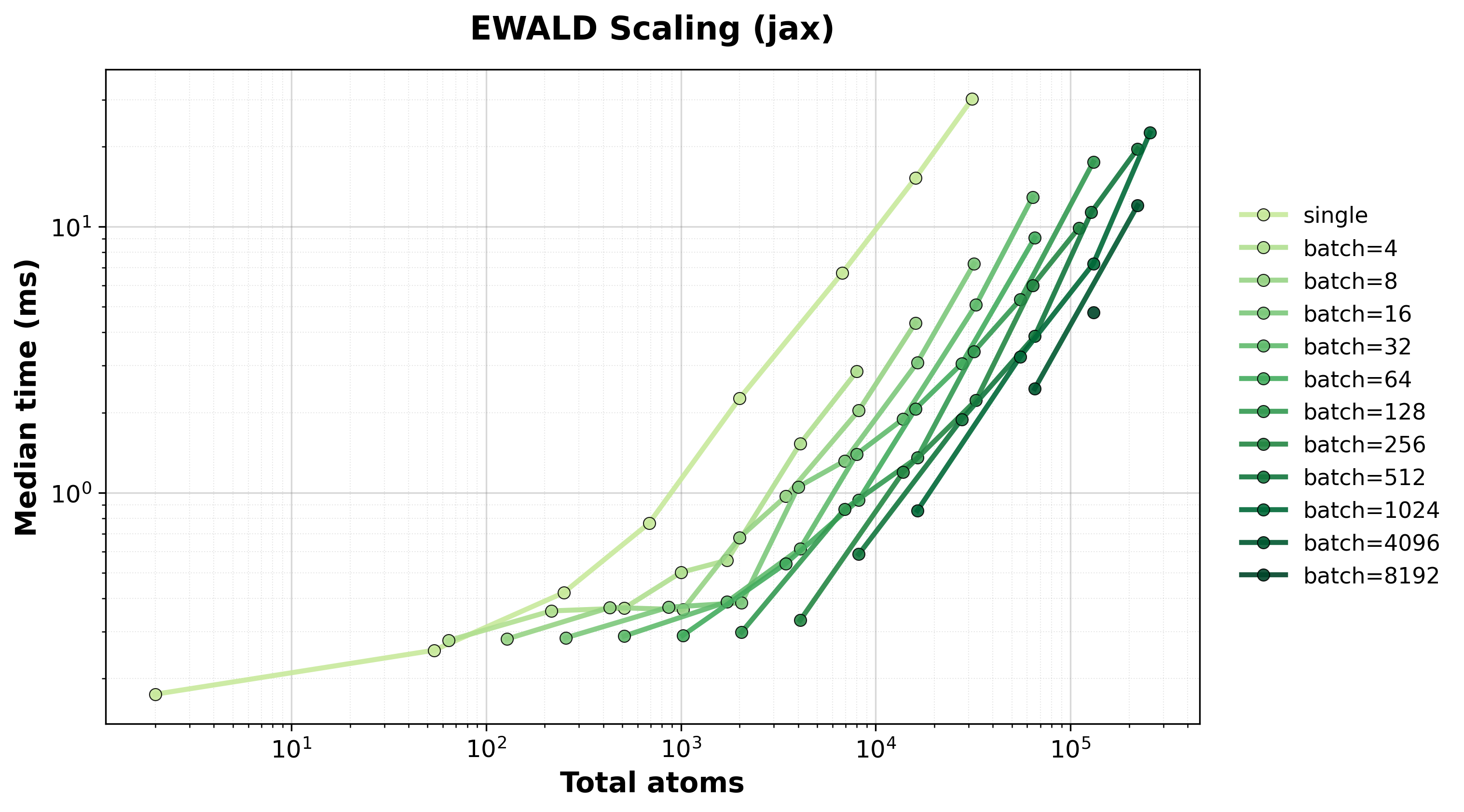
Execution time scaling for single and batched systems.#
Throughput
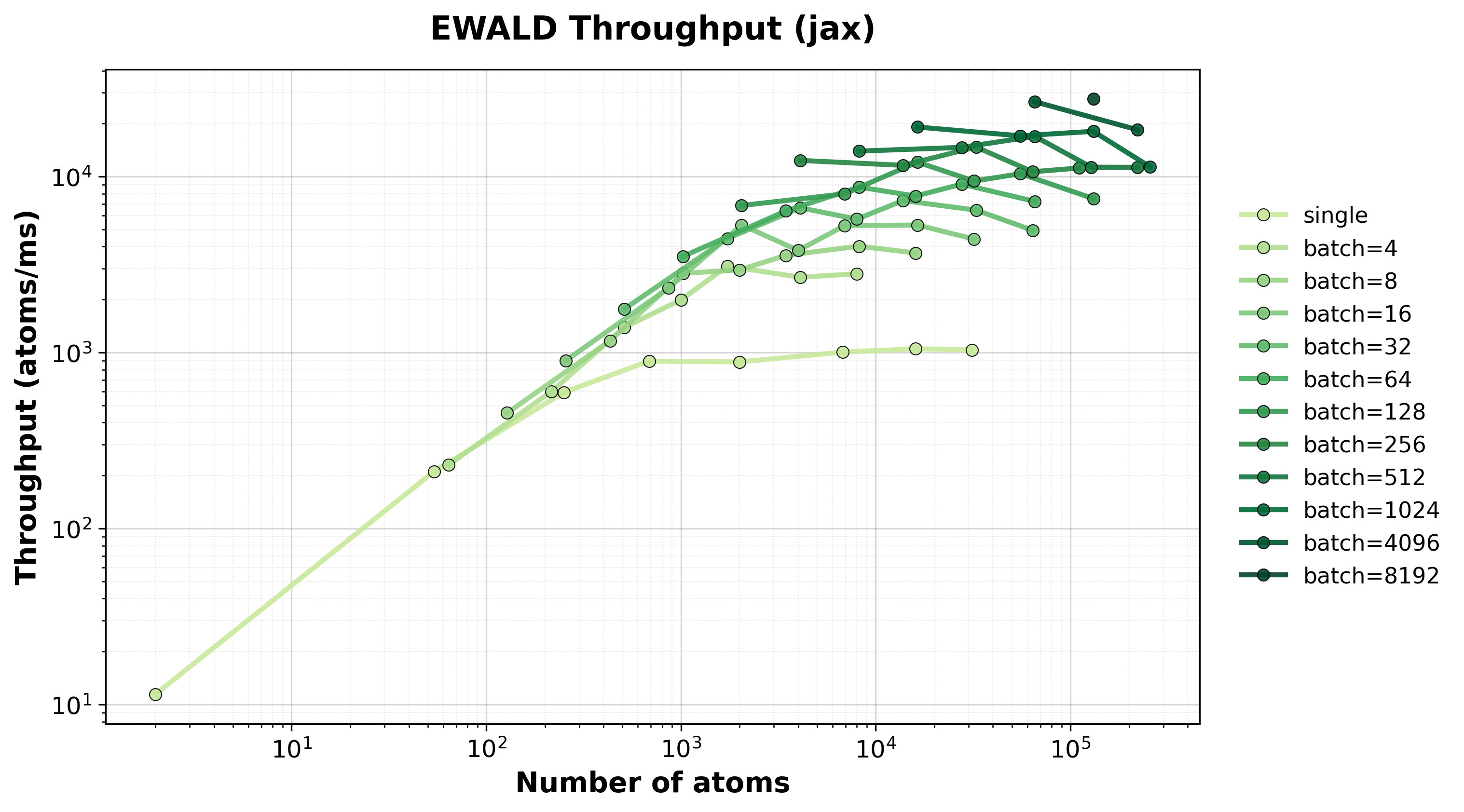
Throughput (atoms/ms) for single and batched systems.#
Memory Usage
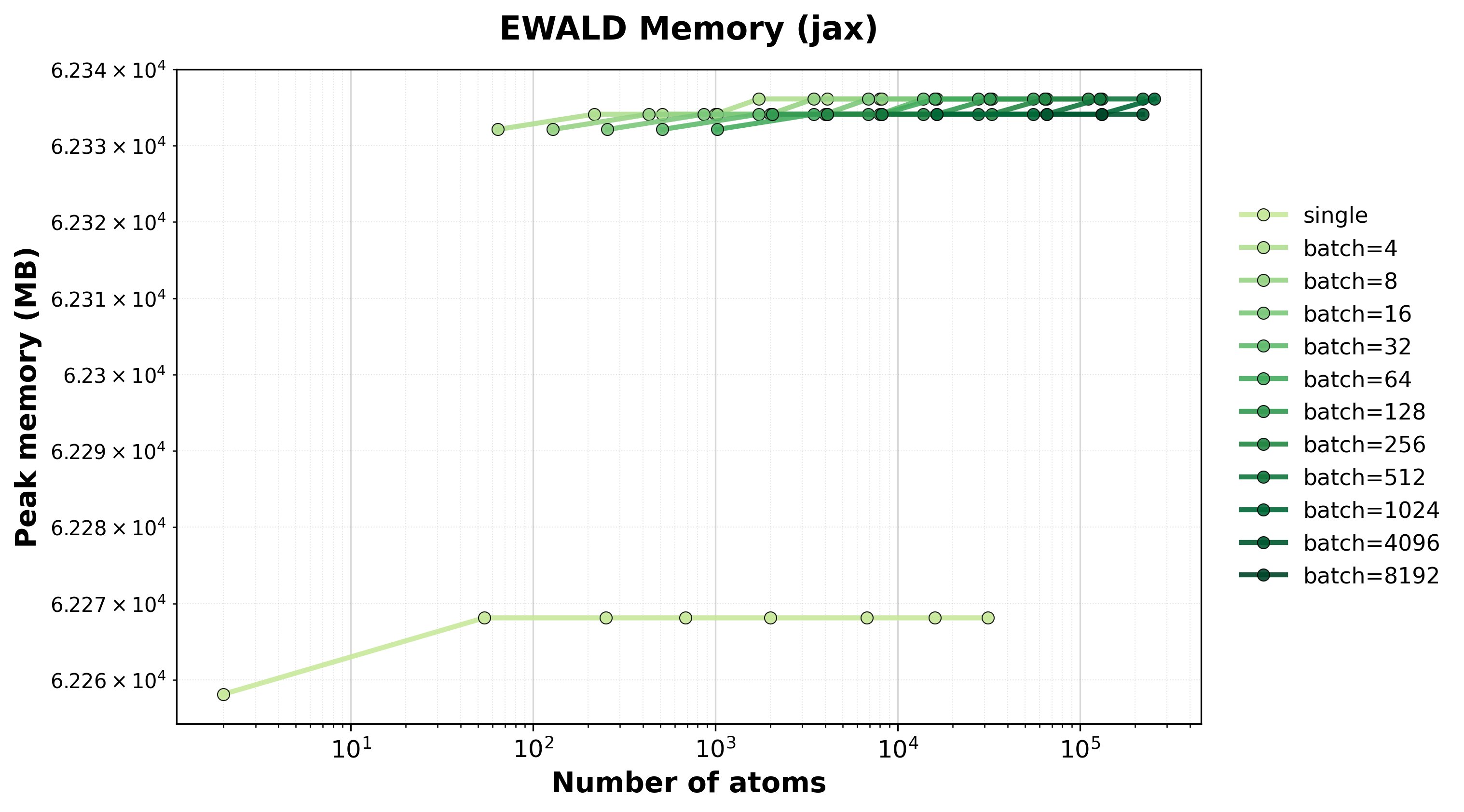
Peak GPU memory consumption for single and batched systems.#
Particle Mesh Ewald (PME)#
PME achieves \(O(N \log N)\) scaling by using FFTs for the reciprocal-space contribution. This is the recommended method for large systems.
Simple comparison of single (non-batched) system computations between backends, where we scale up the size of the supercell.
Time Scaling
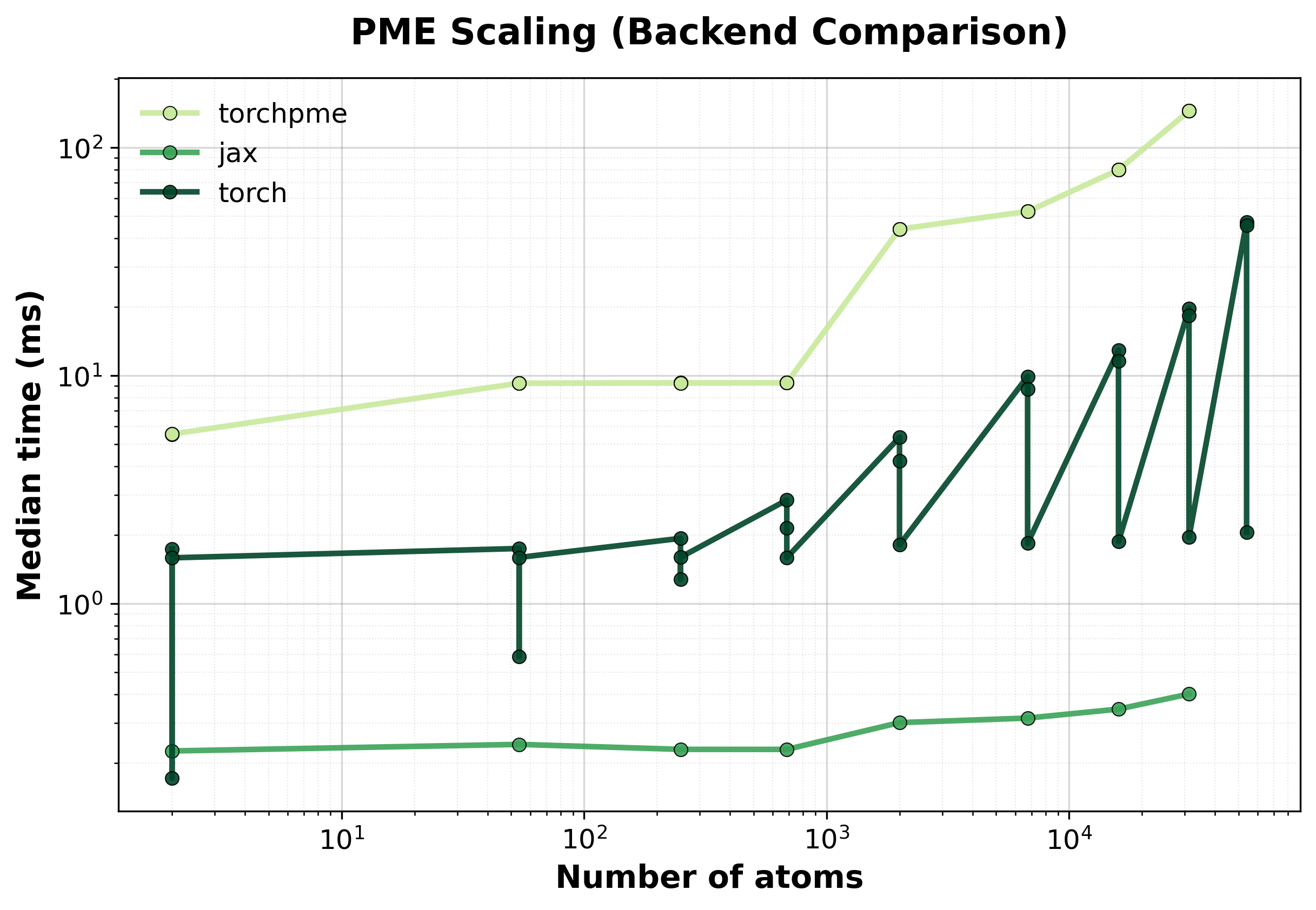
Median execution time comparison between backends for single systems.#
Throughput
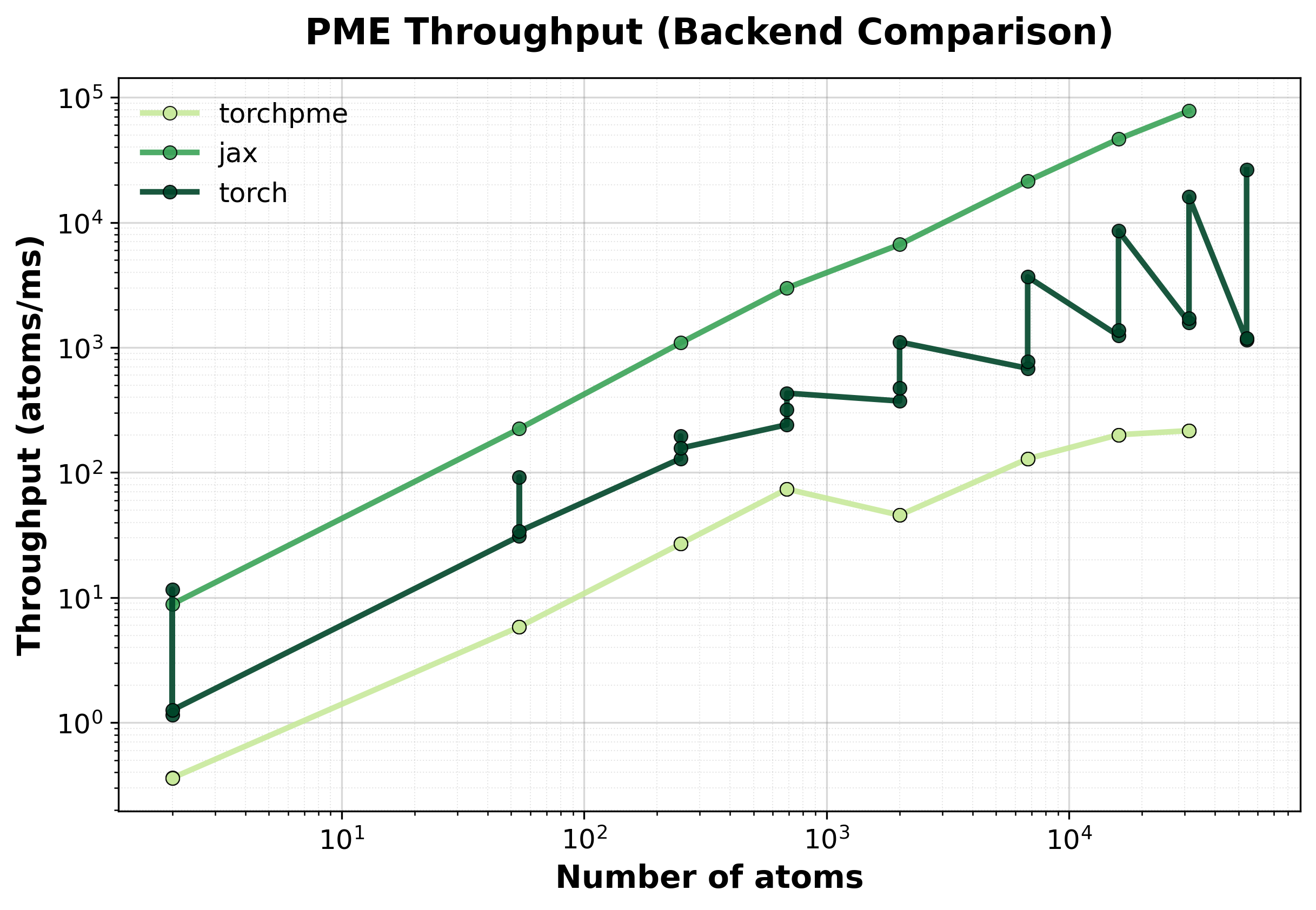
Throughput (atoms/ms) comparison between backends. Higher values indicate better performance.#
Memory Usage
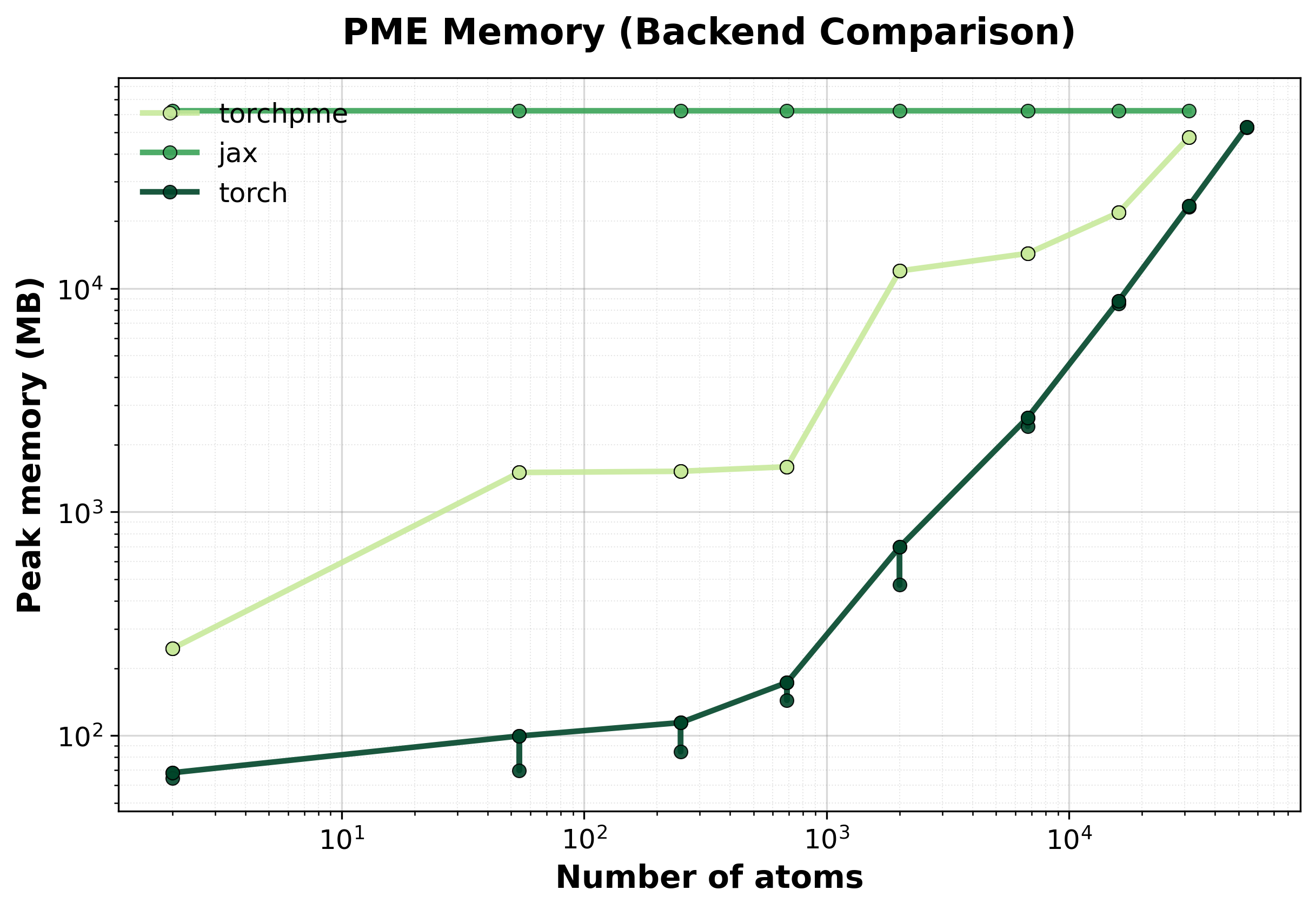
Peak GPU memory consumption comparison between backends. Lower is better, indicating that the backend has lower memory requirements.#
Scaling of single and batched PME computation with the nvalchemiops Torch backend.
Shows how performance scales with different batch sizes.
Time Scaling
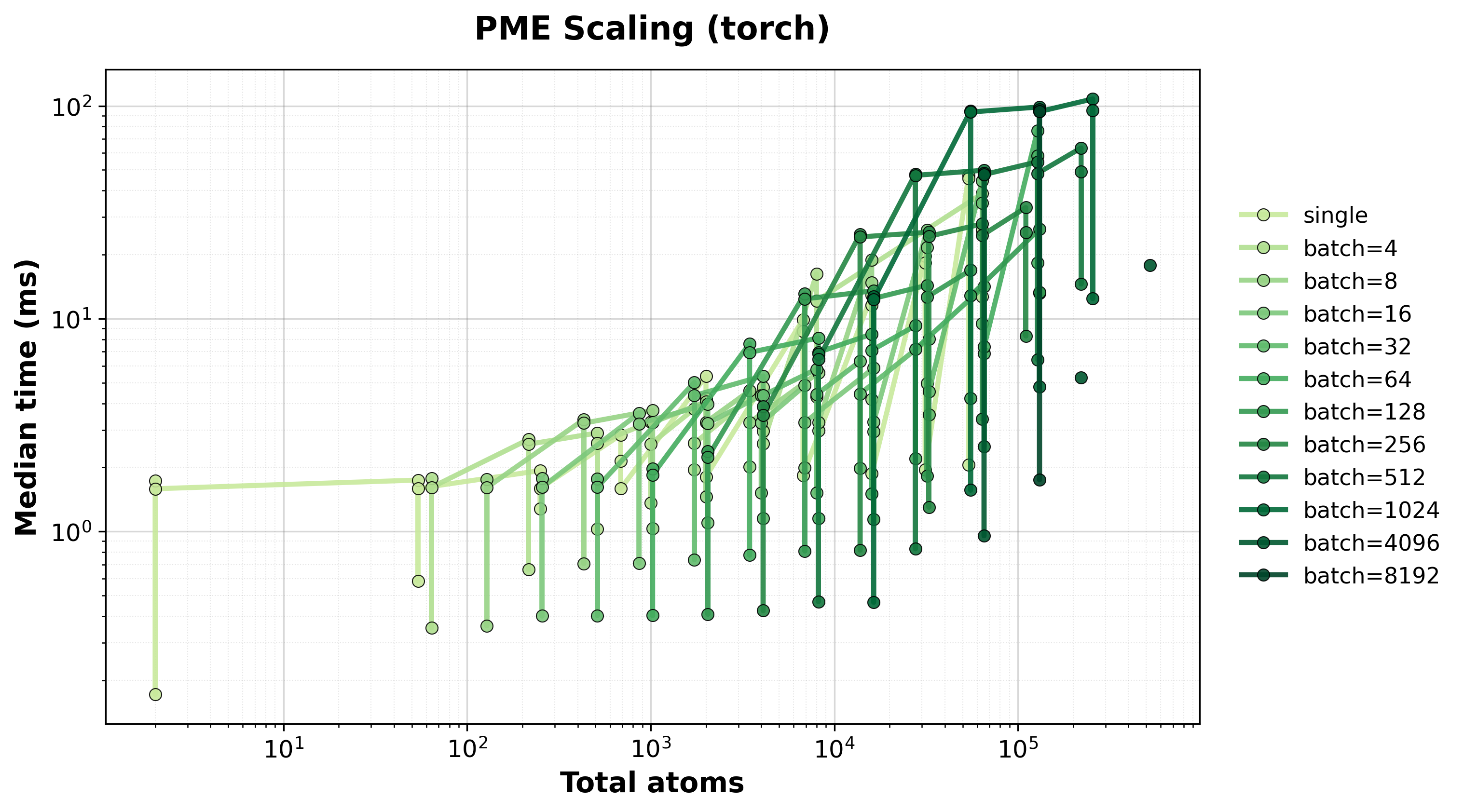
Execution time scaling for single and batched systems.#
Throughput
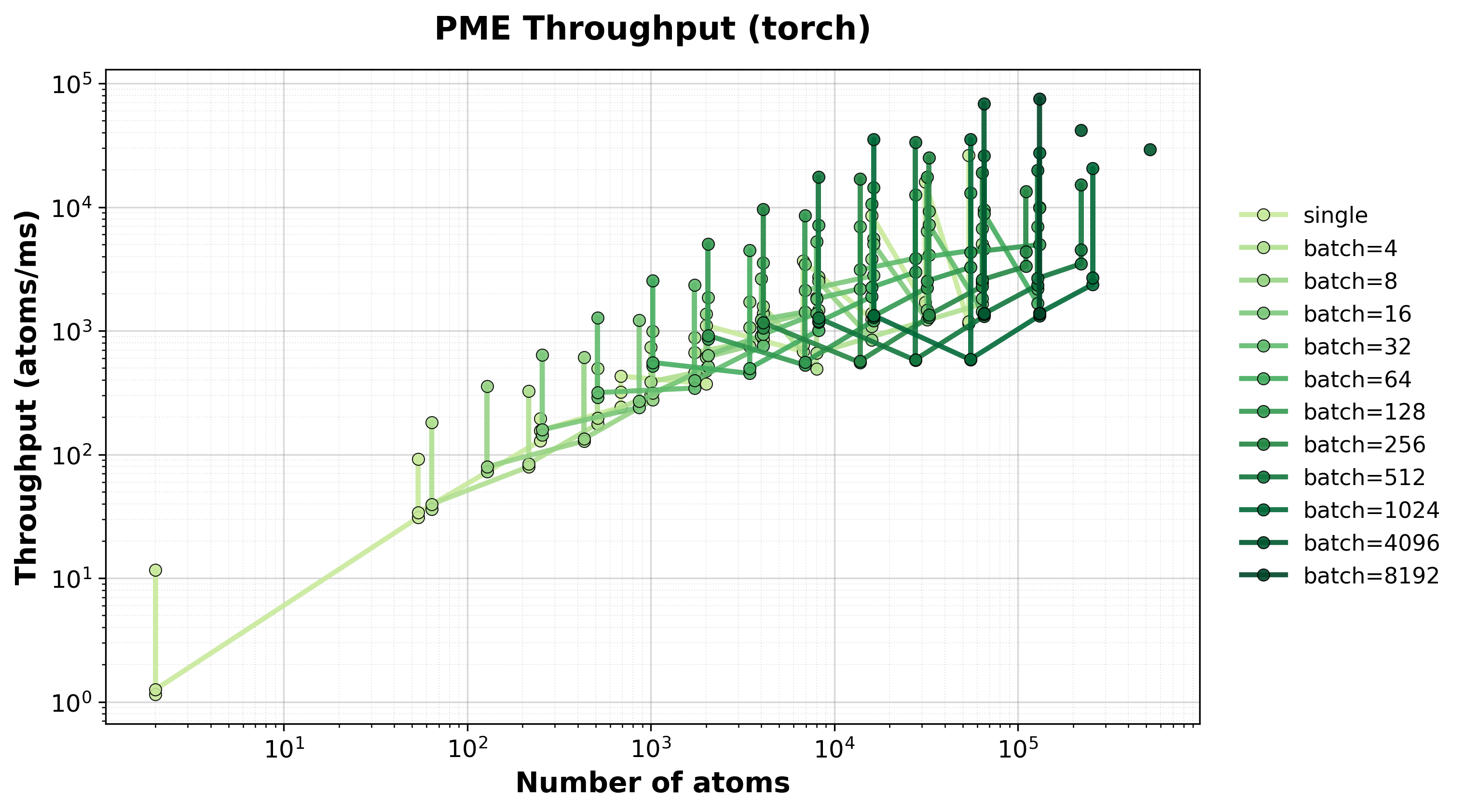
Throughput (atoms/ms) for single and batched systems.#
Memory Usage
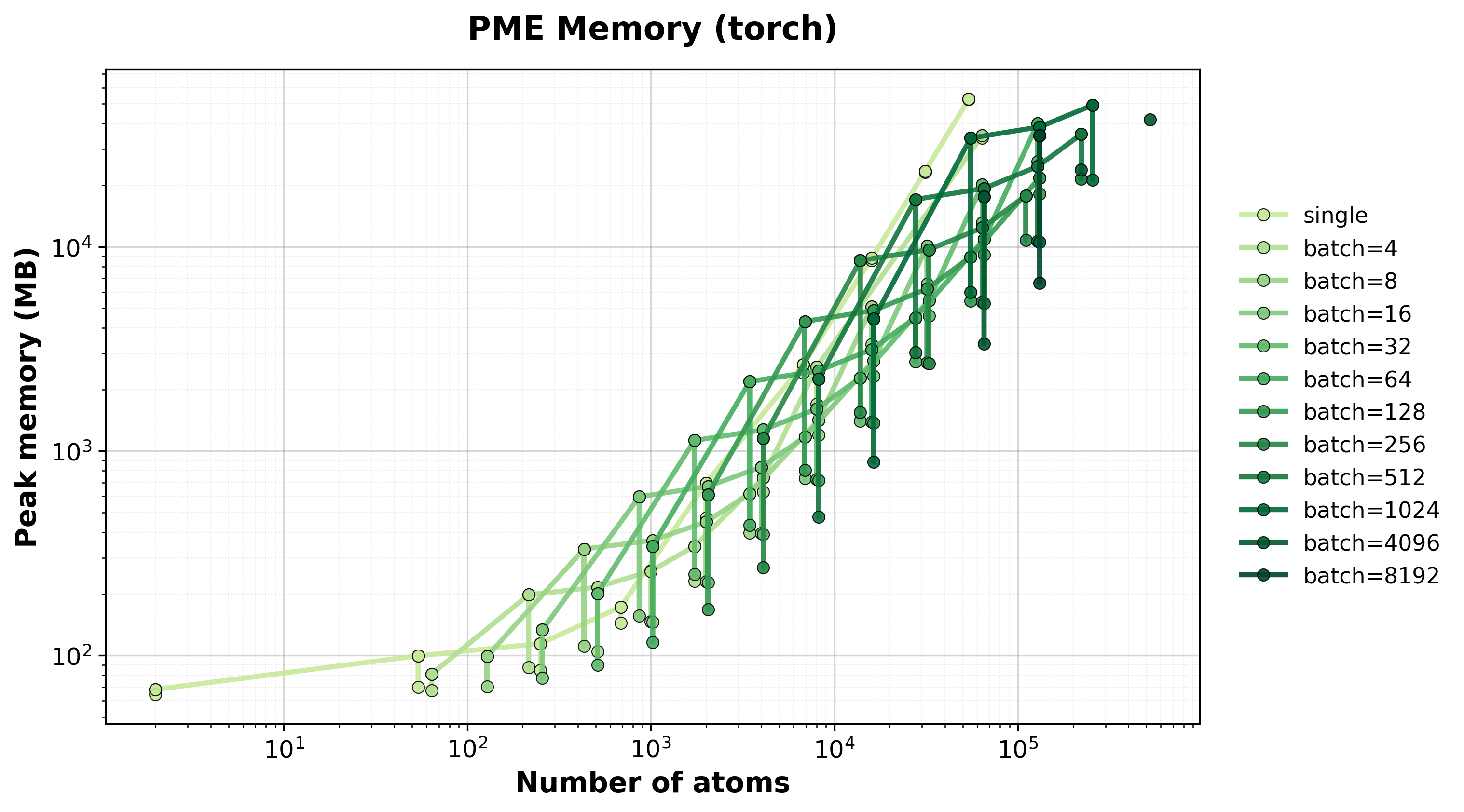
Peak GPU memory consumption for single and batched systems.#
Scaling of single and batched PME computation with the nvalchemiops JAX backend.
Shows how performance scales with different batch sizes.
Time Scaling
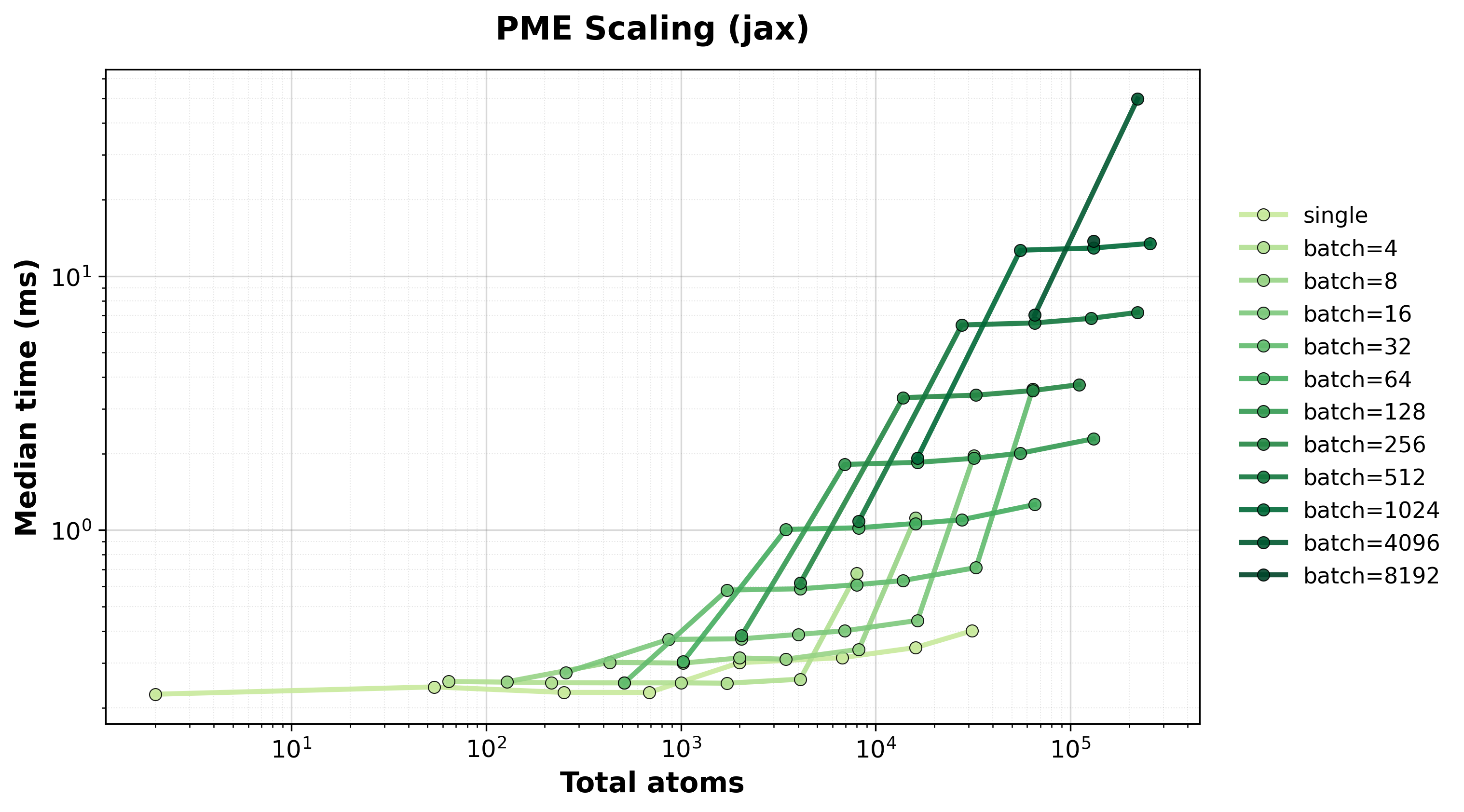
Execution time scaling for single and batched systems.#
Throughput
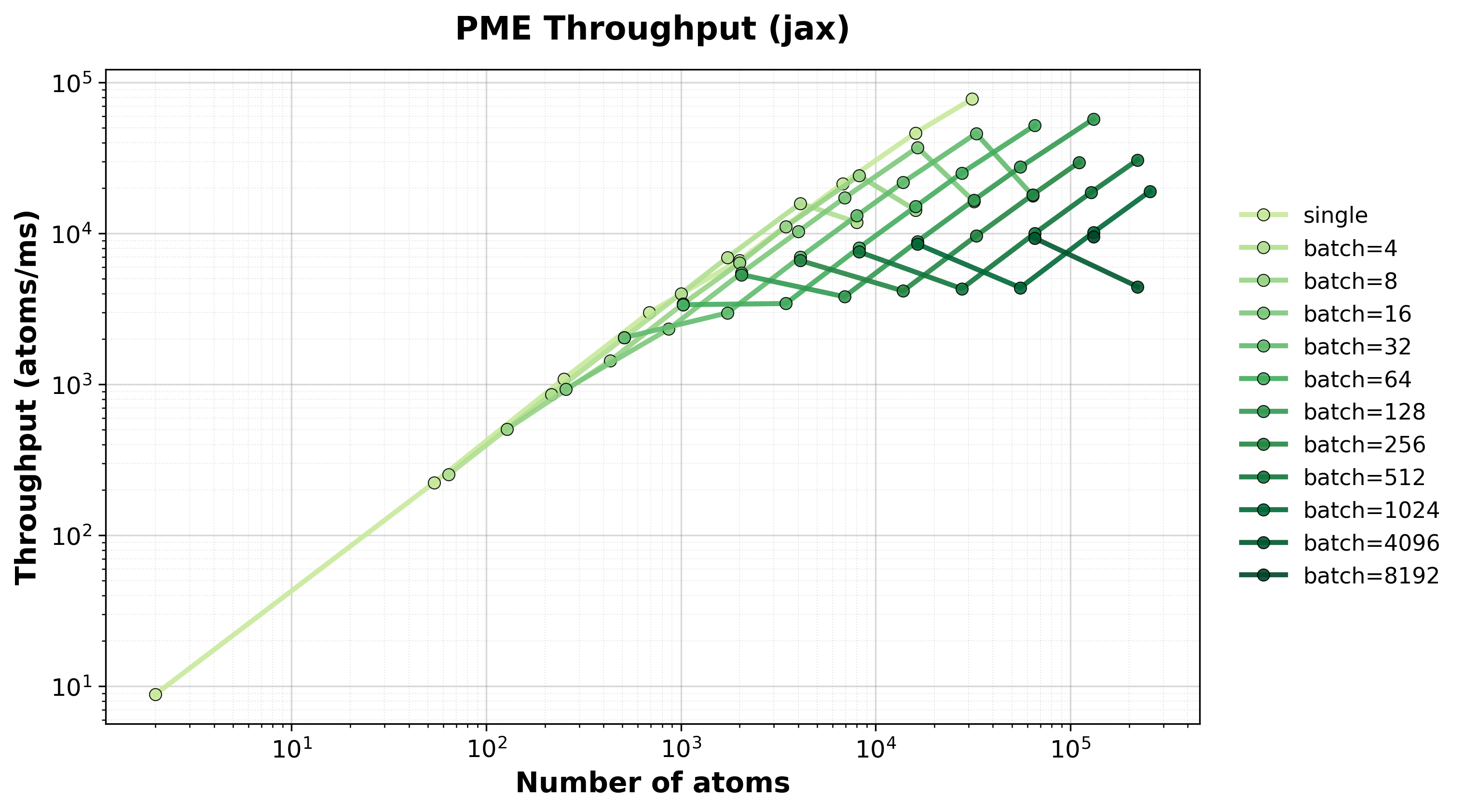
Throughput (atoms/ms) for single and batched systems.#
Memory Usage
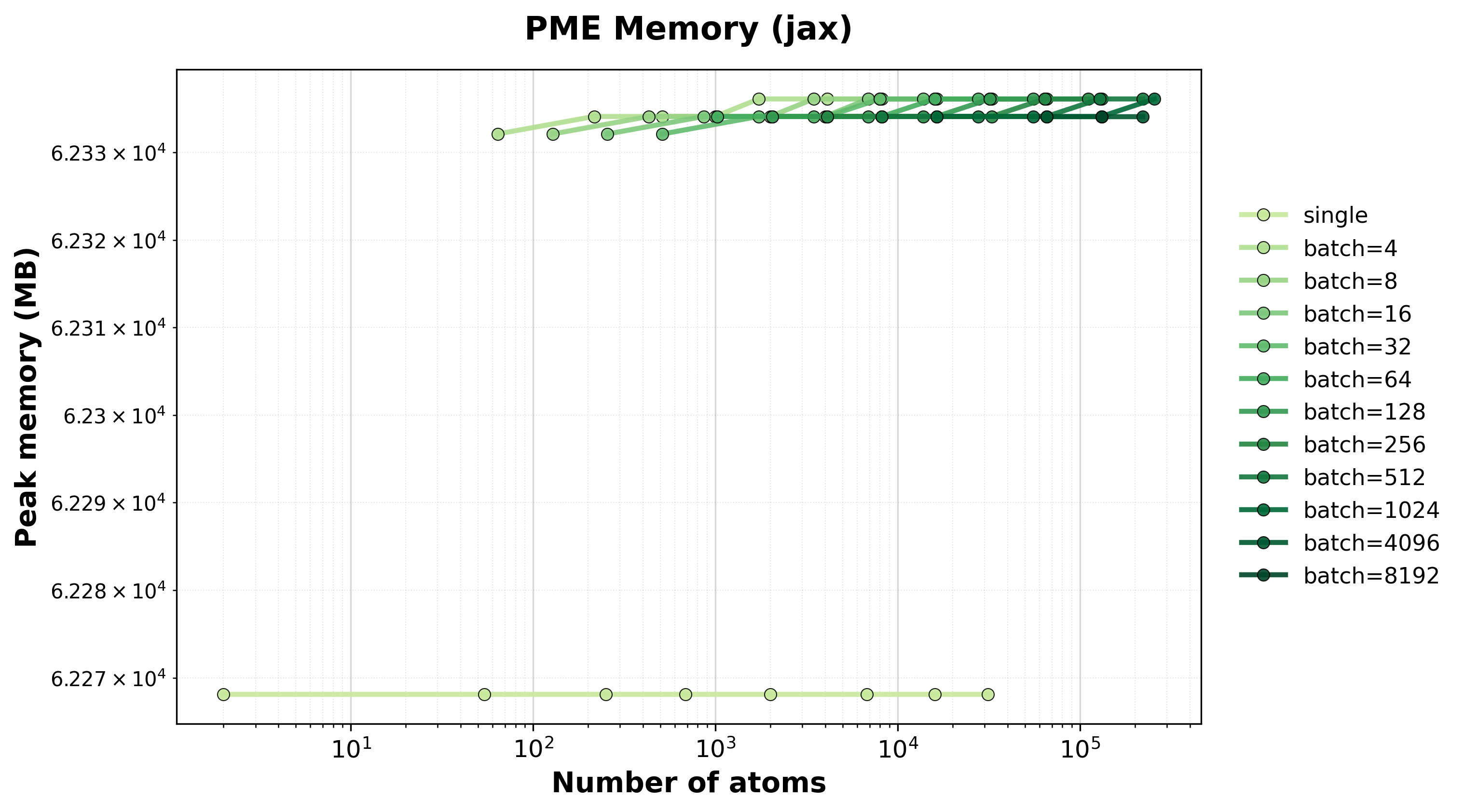
Peak GPU memory consumption for single and batched systems.#
Damped Shifted Force (DSF)#
The DSF method is a purely real-space, pairwise \(O(N)\) electrostatic summation.
Unlike Ewald and PME, DSF has no reciprocal-space component – benchmarks measure
the full calculation directly. DSF supports both CSR neighbor list and dense
neighbor matrix formats, and can be compared against a pure PyTorch reference
implementation (torch_dsf backend).
Note
DSF benchmark plots will be added in a future update. Performance and accuracy
benchmark scripts are available in
benchmarks/interactions/electrostatics/ and can be run on your hardware
using the instructions below.
DSF Parameters#
Parameter |
Value |
|---|---|
|
12.0 |
|
0.2 |
|
|
Hardware Information#
GPU: NVIDIA H100 80GB HBM3
Benchmark Configuration#
Parameter |
Value |
|---|---|
System Type |
FCC crystal lattice with periodic boundaries |
Neighbor List |
Cell list algorithm (\(O(N)\) scaling) |
Warmup Iterations |
3 |
Timing Iterations |
10 |
Precision |
|
Ewald/PME Parameters#
Parameters are automatically estimated using accuracy-based parameter estimation targeting \(10^{-6}\) relative accuracy:
Parameter |
Description |
|---|---|
|
Ewald splitting parameter (auto-estimated) |
|
Reciprocal-space cutoff for Ewald (auto-estimated) |
|
Real-space cutoff distance (auto-estimated) |
|
PME mesh grid size (auto-estimated) |
|
B-spline interpolation order (4) |
Interpreting Results#
total_atoms
: Total number of atoms in the supercell (or across all batched systems).
batch_size
: Number of systems processed simultaneously (1 for single-system mode).
method
: The electrostatics method used (ewald, pme, or dsf).
backend
: The computational backend (torch, jax, torchpme, or torch_dsf).
component
: Which part of the calculation was benchmarked (real, reciprocal, or full).
compute_forces
: Whether forces were computed in addition to energies.
median_time_ms
: Median execution time in milliseconds (lower is better).
peak_memory_mb
: Peak GPU memory usage in megabytes.
Note
Timings include the full electrostatics calculation (real-space + reciprocal-space for “full” mode). Neighbor list construction is excluded from timings.
Running Your Own Benchmarks#
To generate benchmark results for your hardware:
torch Backend (default)#
cd benchmarks/interactions/electrostatics
python benchmark_electrostatics.py \
--config benchmark_config.yaml \
--backend torch \
--method all \
--output-dir ../../../docs/benchmarks/benchmark_results
jax Backend#
cd benchmarks/interactions/electrostatics
python benchmark_electrostatics.py \
--config benchmark_config.yaml \
--backend jax \
--method both \
--output-dir ../../../docs/benchmarks/benchmark_results
torchpme Backend#
cd benchmarks/interactions/electrostatics
python benchmark_electrostatics.py \
--config benchmark_config.yaml \
--backend torchpme \
--method both \
--output-dir ../../../docs/benchmarks/benchmark_results
Options#
--backend {torch,jax,torchpme,torch_dsf,both}
: Computational backend (default: torch). both dispatches per-method:
torch + torchpme for Ewald/PME, torch + torch_dsf for DSF.
--method {ewald,pme,dsf,both,all}
: Electrostatics method (default: both). both = Ewald + PME (backward
compatible). all = Ewald + PME + DSF.
--neighbor-format {list,matrix,both}
: Neighbor format for DSF benchmarks (default: list). Ewald/PME always use matrix.
--dtype {float32,float64}
: Override dtype from config file.
--gpu-sku <name>
: Override GPU SKU name for output files (default: auto-detect).
--config <path>
: Path to YAML configuration file.
DSF-Only Benchmark#
python benchmark_electrostatics.py \
--config benchmark_config.yaml \
--backend both \
--method dsf \
--neighbor-format list
Results will be saved as CSV files and plots will be automatically generated during the next documentation build.
