Dynamics Examples#
This directory contains examples demonstrating the molecular dynamics integrators and geometry optimization algorithms in nvalchemiops.
All examples use realistic Lennard-Jones (LJ) argon systems to demonstrate practical workflows with neighbor list management and periodic boundaries.
MD Integration Examples#
- 01_langevin_integration.py
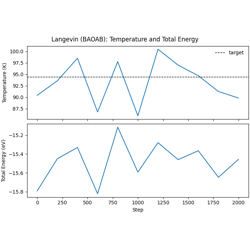
Langevin (BAOAB) dynamics in the NVT ensemble.
Simulates liquid argon near the triple point (94.4 K)
BAOAB splitting for configurational sampling
Temperature monitoring and equilibration
- 02_velocity_verlet_integration.py
Velocity Verlet dynamics in the NVE ensemble.
Symplectic, time-reversible integrator
Energy conservation diagnostics
Stability considerations for LJ systems
- 03_nph_integration.py
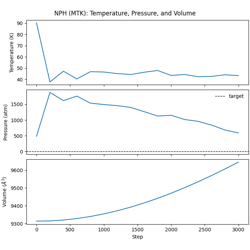
NPH dynamics with MTK barostat.
Constant enthalpy and pressure ensemble
Martyna-Tobias-Klein extended-system barostat
Pressure coupling mechanics
- 04_npt_integration.py
NPT dynamics with MTK barostat + Nosé-Hoover chain thermostat.
Isothermal-isobaric ensemble
Combined temperature and pressure control
Volume fluctuations under constant pressure
- 05_batched_langevin_dynamics.py
Batched Langevin dynamics for multiple independent systems.
Multiple systems packed into single arrays
Per-system temperature targets
Improved GPU utilization for small systems
Geometry Optimization Examples#
- 06_fire_optimization.py
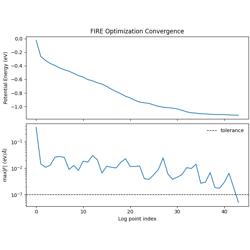
FIRE geometry optimization for a single LJ cluster.
Fast Inertial Relaxation Engine (FIRE) algorithm
Adaptive timestep and velocity mixing
Convergence monitoring (energy, max force)
- 07_fire_variable_cell.py
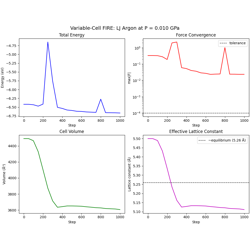
Variable-cell FIRE optimization for joint atom + cell relaxation.
Cell alignment to upper-triangular form
Virial → stress → cell force conversion
Pack/unpack utilities for extended DOF arrays
External pressure equilibration
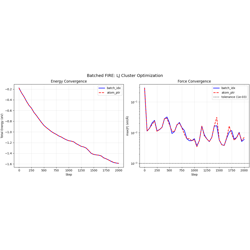
- 08_fire_batched.py
Batched FIRE optimization with two indexing strategies:
batch_idx mode: Each atom tagged with system index
atom_ptr mode (CSR): Atom ranges via pointers
Per-system FIRE parameters adapt independently
Uses
nvalchemiops.batch_utilsfor reductions
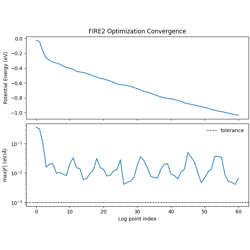
- 09_fire2_optimization.py
FIRE2 geometry optimization for a single LJ cluster.
Improved FIRE variant (Guenole et al., 2020)
Adaptive damping and coupled step/dt scaling
Requires
batch_idxeven for single-system modeHyperparameters are Python scalars (no per-system arrays)
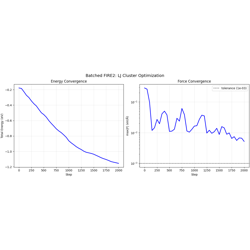
- 10_fire2_batched.py
Batched FIRE2 optimization with
batch_idxbatching.Multiple independent LJ clusters optimized in parallel
Per-system FIRE2 parameters adapt independently
Only supports
batch_idxmode (noatom_ptrvariant)
- 11_fire2_variable_cell.py
Variable-cell FIRE2 optimization for joint atom + cell relaxation.
Same pack/unpack workflow as
07_fire_variable_cell.pyUses
fire2_stepon extended DOF arraysSimpler state than FIRE (no masses, fewer per-system arrays)
Key Concepts#
State Management#
All algorithms use @wp.struct containers for clean organization:
MDState: positions, velocities, forces, massesFIREState: MDState + adaptive parameters (dt, alpha, n_positive)Integration parameters:
VerletParams,LangevinParams,FIREParams
Structs are dtype-agnostic: use wp.vec3f/wp.float32 for single
precision or wp.vec3d/wp.float64 for double precision.
Two-Pass Integrators#
MD integrators use a two-pass design for flexibility:
# Pass 1: Update positions and half-step velocities
velocity_verlet_position_update(state, params)
# User computes forces at new positions
compute_forces(state.positions, state.forces)
# Pass 2: Complete velocity update
velocity_verlet_velocity_finalize(state, state.forces, params)
This allows any force calculation method to be used between the passes.
Batching Modes#
For processing multiple systems efficiently:
batch_idx: Array mapping each atom to its system index. Flexible for heterogeneous systems but requires atomic accumulation.
atom_ptr (CSR): Pointer array where system s owns atoms
[atom_ptr[s], atom_ptr[s+1]). More efficient for homogeneous batches.
Use nvalchemiops.batch_utils for conversions and per-system reductions.
Variable-Cell Optimization#
The cell_filter utilities enable joint optimization of positions and cell:
# 1. Align cell to upper-triangular form
positions, cell = align_cell(positions, cell)
# 2. Pack atomic + cell DOFs into extended arrays
extended_pos = pack_positions_with_cell(positions, cell, ...)
extended_forces = pack_forces_with_cell(forces, cell_force, ...)
# 3. Run optimizer on extended arrays
fire_step(extended_pos, extended_vel, extended_forces, ...)
# 4. Unpack results
positions, cell = unpack_positions_with_cell(extended_pos, ...)
Running the Examples#
Make sure you have warp installed:
pip install warp-lang
Run from the repository root:
python examples/dynamics/01_langevin_integration.py
python examples/dynamics/06_fire_optimization.py
python examples/dynamics/08_fire_batched.py
Or run interactively in VS Code or Jupyter by executing cell-by-cell
(sections marked with # %%).
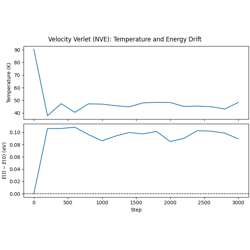
Velocity Verlet Dynamics with Lennard-Jones Potential
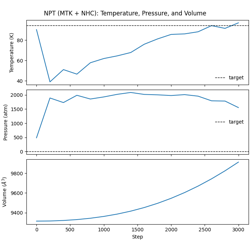
NPT Dynamics (MTK + Nosé-Hoover Chain) with Lennard-Jones Potential
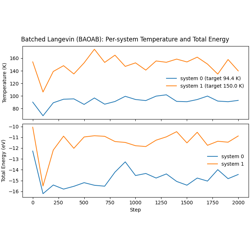
Batched Langevin Dynamics (BAOAB) with Lennard-Jones Potential
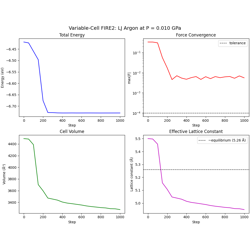
Variable-Cell FIRE2 Optimization with LJ Potential